Implementing in vitro bioactivity data to modernize priority setting of chemical inventories
Main Article Content
Abstract
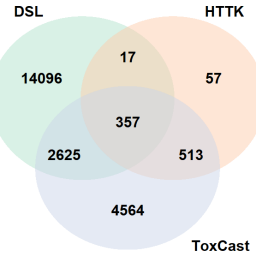
Internationally, there are thousands of existing and newly introduced chemicals in commerce, highlighting the ongoing importance of innovative approaches to identify emerging chemicals of concern. For many chemicals, there is a paucity of hazard and exposure data. Thus, there is a crucial need for efficient and robust approaches to address data gaps and support risk-based prioritization. Several studies have demonstrated the utility of in vitro bioactivity data from the ToxCast program in deriving points of departure (PODs). ToxCast contains data for nearly 1,400 endpoints per chemical, and the bioactivity concentrations, indicative of potential adverse outcomes, can be converted to human-equivalent PODs using high-throughput toxicokinetics (HTTK) modeling. However, data gaps need to be addressed for broader application: the limited chemical space of HTTK and quantitative high-throughput screening data. Here we explore the applicability of in silico models to address these data needs. Specifically, we used ADMET predictor for HTTK predictions and a generalized read-across approach to predict ToxCast bioactivity potency. We applied these models to profile 5,801 chemicals on Canada’s Domestic Substances List (DSL). To evaluate the approach’s performance, bioactivity PODs were compared with in vivo results from the EPA Toxicity Values database for 1,042 DSL chemicals. Comparisons demonstrated that the bioactivity PODs, based on ToxCast data or read-across, were conservative for 95% of the chemicals. Comparing bioactivity PODs to human exposure estimates supports the identification of chemicals of potential interest for further work. The bioactivity workflow shows promise as a powerful screening tool to support effective triaging of chemical inventories.
Article Details

This work is licensed under a Creative Commons Attribution 4.0 International License.
Articles are distributed under the terms of the Creative Commons Attribution 4.0 International license (http://creativecommons.org/licenses/by/4.0/), which permits unrestricted use, distribution and reproduction in any medium, provided the original work is appropriately cited (CC-BY). Copyright on any article in ALTEX is retained by the author(s).
Ankley, G. T., Bennett, R. S., Erickson, R. J. et al. (2010). Adverse outcome pathways: A conceptual framework to support ecotoxicology research and risk assessment. Environ Toxicol Chem 29, 730-741. doi:10.1002/etc.34
Avlasevich, S. L., Bryce, S. M., Cairns, S. E. et al. (2006). In vitro micronucleus scoring by flow cytometry: Differential staining of micronuclei versus apoptotic and necrotic chromatin enhances assay reliability. Environ Mol Mutagen 47, 56-66. doi:10.1002/em.20170
Aylward, L. L. and Hays, S. M. (2011). Consideration of dosimetry in evaluation of ToxCast™ data. J Appl Toxicol 31, 741-751. doi:10.1002/jat.1626
Barter, Z. E., Bayliss, M. K., Beaune, P. H. et al. (2007). Scaling factors for the extrapolation of in vivo metabolic drug clearance from in vitro data: Reaching a consensus on values of human micro-somal protein and hepatocellularity per gram of liver. Curr Drug Metab 8, 33-45. doi:10.2174/138920007779315053
Blackwell, B. R., Ankley, G. T., Corsi, S. R. et al. (2017). An “EAR” on environmental surveillance and monitoring: A case study on the use of exposure-activity ratios (EARs) to prioritize sites, chemicals, and bioactivities of concern in great lakes waters. Environ Sci Technol 51, 8713-8724. doi:10.1021/acs.est.7b01613
Bryce, S. M., Avlasevich, S. L., Bemis, J. C. et al. (2010). Miniaturized flow cytometric in vitro micronucleus assay represents an efficient tool for comprehensively characterizing genotoxicity dose-response relationships. Mutat Res 703, 191-199. doi:10.1016/j.mrgentox.2010.08.020
Bryce, S. M., Bernacki, D. T., Smith-Roe, S. L. et al. (2018). Investigating the generalizability of the MultiFlow® DNA damage assay and several companion machine learning models with a set of 103 diverse test chemicals. Toxicol Sci 162, 146-166. doi:10.1093/toxsci/kfx235
Cao, Y., Charisi, A., Cheng, L. et al. (2008). ChemmineR: A compound mining framework for R. Bioinformatics 24, 1733-1734. doi:10.1093/bioinformatics/btn307
Chao, C. and Engelward, B. P. (2020). Applications of CometChip for environmental health studies. Chem Res Toxicol 33, 1528-1538. doi:10.1021/acs.chemrestox.9b00393
Cohen Hubal, E. A., Richard, A., Aylward, L. et al. (2010). Advancing exposure characterization for chemical evaluation and risk assessment. J Toxicol Environ Health B 13, 299-313. doi:10.1080/10937404.2010.483947
Corsi, S. R., De Cicco, L. A., Villeneuve, D. L. et al. (2019). Prioritizing chemicals of ecological concern in great lakes tributaries using high-throughput screening data and adverse outcome pathways. Sci Total Environ 686, 995-1009. doi:10.1016/j.scitotenv.2019.05.457
Dennis, K. K., Marder, E., Balshaw, D. M. et al. (2017). Biomonitoring in the era of the exposome. Environ Health Perspect 125, 502-510. doi:10.1289/EHP474
EFSA and WHO – European Food Safety Authority and World Health Organization (2016). Review of the threshold of toxicological concern (TTC) approach and development of new TTC decision tree. EFSA Supporting Publications 13, 1006E. doi:10.2903/sp.efsa.2016.EN-1006
Fan, X., Katuri, G. P., Caza, A. A. et al. (2021). Simultaneous measurement of 16 bisphenol A analogues in house dust and evaluation of two sampling techniques. Emerging Contaminants 7, 1-9. doi:10.1016/j.emcon.2020.12.001
Farmahin, R., Williams, A., Kuo, B. et al. (2017). Recommended approaches in the application of toxicogenomics to derive points of departure for chemical risk assessment. Arch Toxicol 91, 2045-2065. doi:10.1007/s00204-016-1886-5
Filer, D. L., Kothiya, P., Setzer, R. W. et al. (2016). tcpl: The ToxCast pipeline for high-throughput screening data. Bioinformatics 33, 618-620. doi:10.1093/bioinformatics/btw680
Gannon, A. M., Moreau, M., Farmahin, R. et al. (2019). Hexabromocyclododecane (HBCD): A case study applying tiered testing for human health risk assessment. Food Chem Toxicol 131, 110581. doi:10.1016/j.fct.2019.110581
Harrill, J., Shah, I., Setzer, R. W. et al. (2019). Considerations for strategic use of high-throughput transcriptomics chemical screening data in regulatory decisions. Curr Opin Toxicol 15, 64-75. doi:10.1016/j.cotox.2019.05.004
Harrill, J. A., Everett, L. J., Haggard, D. E. et al. (2021). High-throughput transcriptomics platform for screening environmental chemicals. Toxicol Sci 181, 68-89. doi:10.1093/toxsci/kfab009
Health Canada (2016). Science Approach Document: Threshold of Toxicological Concern (TTC)-Based Approach for Certain Substances (existing substance risk assessment bureau, ed.). Government of Canada, Ottawa, Ontario, Canada. https://bit.ly/3I8p2Ho
Health Canada (2021). Science approach document – Bioactivity exposure ratio: Application in priority setting and risk assessment. Canada Gazette 155. https://www.canada.ca/content/dam/eccc/documents/pdf/pded/bioactivity-exposure-ratio/Science-approach-document-bioactivity-exposure-ratio.pdf
Helman, G., Patlewicz, G. and Shah, I. (2019). Quantitative prediction of repeat dose toxicity values using GenRA. Regul Toxicol Pharmacol 109, 104480. doi:10.1016/j.yrtph.2019.104480
Hendriks, G., Atallah, M., Morolli, B. et al. (2012). The ToxTracker assay: Novel GFP reporter systems that provide mechanistic insight into the genotoxic properties of chemicals. Toxicol Sci 125, 285-298. doi:10.1093/toxsci/kfr281
Hsieh, J., Smith-Roe, S. L., Huang, R. et al. (2019). Identifying compounds with genotoxicity potential using Tox21 high-throughput screening assays. Chem Res Toxicol 32, 1384-1401. doi:10.1021/acs.chemrestox.9b00053
Johnson, C. L., Dohrmann, S. M., Burt, V. L. et al. (2014). National health and nutrition examination survey: Sample design, 2011-2014. Vital Health Stat 2, 1-33.
Judson, R. S., Kavlock, R. J., Setzer, R. W. et al. (2011). Estimating toxicity-related biological pathway altering doses for high-throughput chemical risk assessment. Chem Res Toxicol 24, 451-462. doi:10.1021/tx100428e
Kavlock, R. J., Bahadori, T., Barton-Maclaren, T. S. et al. (2018). Accelerating the pace of chemical risk assessment. Chem Res Toxicol 31, 287-290. doi:10.1021/acs.chemrestox.7b00339
Kroes, R., Renwick, A., Cheeseman, M. et al. (2004). Structure-based thresholds of toxicological concern (TTC): Guidance for application to substances present at low levels in the diet. Food Chem Toxicol 42, 65-83. doi:10.1016/j.fct.2003.08.006
Li, H., Hyduke, D. R., Chen, R. et al. (2015). Development of a toxicogenomics signature for genotoxicity using a dose‐optimization and informatics strategy in human cells. Environ Mol Mutagen 56, 505-519. doi:10.1002/em.21941
Li, H. H., Chen, R., Hyduke, D. R. et al. (2017). Development and validation of a high-throughput transcriptomic biomarker to address 21st century genetic toxicology needs. Proc Natl Acad Sci USA 114, E10881-E10889. doi:10.1073/pnas.1714109114
Lipinski, C. A., Lombardo, F., Dominy, B. W. et al. (1997). Experimental and computational approaches to estimate solubility and permeability in drug discovery and development settings. Adv Drug Deliv Rev 23, 3-25. doi:10.1016/s0169-409x(00)00129-0
Maertens, R. M., Long, A. S. and White, P. A. (2017). Performance of the in vitro transgene mutation assay in MutaMouse FE1 cells: Evaluation of nine misleading (“false”) positive chemicals. Environ Mol Mutagen 58, 582-591. doi:10.1002/em.22125
Mansouri, K., Abdelaziz, A., Rybacka, A. et al. (2016). CERAPP: Collaborative estrogen receptor activity prediction project. Environ Health Perspect 124, 1023-1033. doi:10.1289/ehp.1510267
Mansouri, K., Kleinstreuer, N., Abdelaziz, A. M. et al. (2020). CoMPARA: Collaborative modeling project for androgen receptor activity. Environ Health Perspect 128, 027002. doi:10.1289/EHP5580
O’Boyle, N. M., Banck, M., James, C. A. et al. (2011). Open Babel: An open chemical toolbox. J Cheminform 3, 33. doi:10.1186/1758-2946-3-33
OECD (2019). International Best Practices for Identification of Priorities within Chemicals Management Systems. OECD Series on Testing and Assessment, No. 314. OECD Publishing, Paris. https://doi.org/10.1787/0fafd6f5-en
Patlewicz, G., Jeliazkova, N., Safford, R. et al. (2008). An evaluation of the implementation of the Cramer classification scheme in the Toxtree software. SAR QSAR Environ Res 19, 495-524. doi:10.1080/10629360802083871
Patlewicz, G., Kuseva, C., Kesova, A. et al. (2014). Towards AOP application – Implementation of an integrated approach to testing and assessment (IATA) into a pipeline tool for skin sensitization. Regul Toxicol Pharmacol 69, 529-545. doi:10.1016/j.yrtph.2014.06.001
Patlewicz, G., Wambaugh, J. F., Felter, S. P. et al. (2018). Utilizing threshold of toxicological concern (TTC) with high throughput exposure predictions (HTE) as a risk-based prioritization approach for thousands of chemicals. Comput Toxicol 7, 58-67. doi:10.1016/j.comtox.2018.07.002
Paul Friedman, K., Papineni, S., Marty, M. S. et al. (2016). A predictive data-driven framework for endocrine prioritization: A triazole fungicide case study. Crit Rev Toxicol 46, 785-833. doi:10.1080/10408444.2016.1193722
Paul Friedman, K., Gagne, M., Loo, L. et al. (2019). Utility of in vitro bioactivity as a lower bound estimate of in vivo adverse effect levels and in risk-based prioritization. Toxicol Sci 173, 202-225. doi:10.1093/toxsci/kfz201
Pearce, R. G., Setzer, R. W., Strope, C. L. et al. (2017). httk: R package for high-throughput toxicokinetics. J Stat Softw 79, 1-26. doi:10.18637/jss.v079.i04
Phillips, K. A., Wambaugh, J. F., Grulke, C. M. et al. (2017). High-throughput screening of chemicals as functional substitutes using structure-based classification models. Green Chem 19, 1063-1074. doi:10.1039/C6GC02744J
Pourchet, M., Debrauwer, L., Klanova, J. et al. (2020). Suspect and non-targeted screening of chemicals of emerging concern for human biomonitoring, environmental health studies and support to risk assessment: From promises to challenges and harmonisation issues. Environ Int 139, 105545. doi:10.1016/j.envint.2020.105545
Pradeep, P., Patlewicz, G., Pearce, R. et al. (2020). Using chemical structure information to develop predictive models for in vitro toxicokinetic parameters to inform high-throughput risk-assessment. Comput Toxicol 16, 100136. doi:10.1016/j.comtox.2020.100136
Pradeep, P., Judson, R., DeMarini, D. M. et al. (2021). An evaluation of existing QSAR models and structural alerts and development of new ensemble models for genotoxicity using a newly compiled experimental dataset. Comput Toxicol 18, 100167. doi:10.1016/j.comtox.2021.100167
Rager, J. E., Strynar, M. J., Liang, S. et al. (2016). Linking high resolution mass spectrometry data with exposure and toxicity forecasts to advance high-throughput environmental monitoring. Environ Int 88, 269-280. doi:10.1016/j.envint.2015.12.008
Rajkumar, A., Luu, T., Beal, M. A. et al. (2021). Elucidation of the effects of bisphenol A and structural analogs on germ and steroidogenic cells using single cell high-content imaging. Toxicol Sci 180, 224-238. doi:10.1093/toxsci/kfab012
Richard, A. M., Judson, R. S., Houck, K. A. et al. (2016). ToxCast chemical landscape: Paving the road to 21st century toxicology. Chem Res Toxicol 29, 1225-1251. doi:10.1021/acs.chemrestox.6b00135
Ring, C. L., Pearce, R. G., Setzer, R. W. et al. (2017). Identifying populations sensitive to environmental chemicals by simulating toxicokinetic variability. Environ Int 106, 105-118. doi:10.1016/j.envint.2017.06.004
Salk, J. J., Schmitt, M. W. and Loeb, L. A. (2018). Enhancing the accuracy of next-generation sequencing for detecting rare and subclonal mutations. Nat Rev Genet 19, 269. doi:10.1038/nrg.2017.117
Salk, J. J. and Kennedy, S. R. (2020). Next‐generation genotoxicology: Using modern sequencing technologies to assess somatic mutagenesis and cancer risk. Environ Mol Mutagen 61, 135-151. doi:10.1002/em.22342
Schmitt, M. W., Kennedy, S. R., Salk, J. J. et al. (2012). Detection of ultra-rare mutations by next-generation sequencing. Proc Natl Acad Sci U S A 109, 14508-14513. doi:10.1073/pnas.1208715109
Shah, I., Liu, J., Judson, R. S. et al. (2016). Systematically evaluating read-across prediction and performance using a local validity approach characterized by chemical structure and bioactivity information. Regul Toxicol Pharmacol 79, 12-24. doi:10.1016/j.yrtph.2016.05.008
Sipes, N. S., Martin, M. T., Kothiya, P. et al. (2013). Profiling 976 ToxCast chemicals across 331 enzymatic and receptor signaling assays. Chem Res Toxicol 26, 878-895. doi:10.1021/tx400021f
Sipes, N. S., Wambaugh, J. F., Pearce, R. et al. (2017). An intuitive approach for predicting potential human health risk with the Tox21 10k library. Environ Sci Technol 51, 10786-10796. doi:10.1021/acs.est.7b00650
Steinbeck, C., Han, Y., Kuhn, S. et al. (2003). The chemistry development kit (CDK): An open-source java library for chemo-and bioinformatics. J Chem Inf Comput Sci 43, 493-500. doi:10.1021/ci025584y
Thomas, R. S., Philbert, M. A., Auerbach, S. S. et al. (2013a). Incorporating new technologies into toxicity testing and risk assessment: Moving from 21st century vision to a data-driven framework. Toxicol Sci 136, 4-18. doi:10.1093/toxsci/kft178
Thomas, R. S., Wesselkamper, S. C., Wang, N. C. Y. et al. (2013b). Temporal concordance between apical and transcriptional points of departure for chemical risk assessment. Toxicol Sci 134, 180-194. doi:10.1093/toxsci/kft094
Thomas, R. S., Bahadori, T., Buckley, T. J. et al. (2019). The next generation blueprint of computational toxicology at the US environmental protection agency. Toxicol Sci 169, 317-332. doi:10.1093/toxsci/kfz058
Tilley, S. K., Reif, D. M. and Fry, R. C. (2017). Incorporating ToxCast and Tox21 datasets to rank biological activity of chemicals at superfund sites in North Carolina. Environ Int 101, 19-26. doi:10.1016/j.envint.2016.10.006
Tonnelier, A., Coecke, S. and Zaldívar, J. (2012). Screening of chemicals for human bioaccumulative potential with a physiologically based toxicokinetic model. Arch Toxicol 86, 393-403. doi:10.1007/s00204-011-0768-0
Turley, A. E., Isaacs, K. K., Wetmore, B. A. et al. (2019). Incorporating new approach methodologies in toxicity testing and exposure assessment for tiered risk assessment using the RISK21 approach: Case studies on food contact chemicals. Food Chem Toxicol 134, 110819. doi:10.1016/j.fct.2019.110819
US EPA – US Environmental Protection Agency (2015). ToxCast & Tox21 MySQL database invitrodb (version 3).
Villeneuve, D. L., Crump, D., Garcia-Reyero, N. et al. (2014). Adverse outcome pathway (AOP) development I: Strategies and principles. Toxicol Sci 142, 312-320. doi:10.1093/toxsci/kfu199
Wambaugh, J. F., Setzer, R. W., Reif, D. M. et al. (2013). High-throughput models for exposure-based chemical prioritization in the ExpoCast project. Environ Sci Technol 47, 8479-8488. doi:10.1021/es400482g
Wambaugh, J. F., Wetmore, B. A., Pearce, R. et al. (2015). Toxicokinetic triage for environmental chemicals. Toxicol Sci 147, 55-67. doi:10.1093/toxsci/kfv118
Wambaugh, J. F., Hughes, M. F., Ring, C. L. et al. (2018). Evaluating in vitro-in vivo extrapolation of toxicokinetics. Toxicol Sci 163, 152-169. doi:10.1093/toxsci/kfy020
Weingeist, D. M., Ge, J., Wood, D. K. et al. (2013). Single-cell microarray enables high-throughput evaluation of DNA double-strand breaks and DNA repair inhibitors. Cell Cycle 12, 907-915. doi:10.4161/cc.23880
Wetmore, B. A., Wambaugh, J. F., Ferguson, S. S. et al. (2011). Integration of dosimetry, exposure, and high-throughput screening data in chemical toxicity assessment. Toxicol Sci 125, 157-174. doi:10.1093/toxsci/kfr254
Wetmore, B. A., Wambaugh, J. F., Ferguson, S. S. et al. (2013). Relative impact of incorporating pharmacokinetics on predicting in vivo hazard and mode of action from high-throughput in vitro toxicity assays. Toxicol Sci 132, 327-346. doi:10.1093/toxsci/kft012
Wetmore, B. A. (2015). Quantitative in vitro-to-in vivo extrapolation in a high-throughput environment. Toxicology 332, 94-101. doi:10.1016/j.tox.2014.05.012
White, P. A., Douglas, G. R., Gingerich, J. et al. (2003). Development and characterization of a stable epithelial cell line from muta™ mouse lung. Environ Mol Mutagen 42, 166-184. doi:10.1002/em.10185
Wignall, J. A., Muratov, E., Sedykh, A. et al. (2018). Conditional toxicity value (CTV) predictor: An in silico approach for generating quantitative risk estimates for chemicals. Environ Health Perspect 126, 057008. doi:10.1289/EHP2998
Williams, A. J., Grulke, C. M., Edwards, J. et al. (2017). The CompTox chemistry dashboard: A community data resource for environmental chemistry. J Cheminform 9, 61. doi:10.1186/s13321-017-0247-6
Yang, C., Tarkhov, A., Marusczyk, J. et al. (2015). New publicly available chemical query language, CSRML, to support chemotype representations for application to data mining and modeling. J Chem Inf Model 55, 510-528. doi:10.1021/ci500667v